University of Bath
Research Computing Team (DDaT)
Anatra HPC Documentation
Running Containerized Jobs with Apptainer¶
What is Apptainer?¶
Apptainer (formerly Singularity) is a container platform designed for HPC environments. It allows you to package software, libraries, and dependencies into a single container file, ensuring your analysis runs consistently across different systems. Unlike Docker, Apptainer doesn't require root privileges to run containers, making it ideal for shared HPC systems like Anatra.
Getting Started with Apptainer¶
Loading the Module¶
Before using Apptainer, load the module:
module load apptainer/1.4.1
# Verify it's loaded
apptainer --version
Building Your First Container¶
Since building containers requires elevated privileges, you have two options on Anatra:
Option 1: Remote Build (Recommended)
Create a definition file biotools.def:
Bootstrap: docker
From: condaforge/mambaforge:latest
%post
# Configure conda channels in the correct priority order
conda config --add channels defaults
conda config --add channels bioconda
conda config --add channels conda-forge
conda config --set channel_priority strict
# Install mamba for faster dependency resolution
conda install -n base -c conda-forge mamba -y
# Create environment with your bioinformatics tools
mamba create -n biotools python=3.10 numpy scipy -y
# Clean up to reduce container size
conda clean --all -y
%environment
# Make tools available in PATH
export PATH=/opt/conda/envs/biotools/bin:$PATH
%labels
Author username@bath.ac.uk
Version 1.0
Description Bioinformatics tools via Conda
%runscript
echo "Bioinformatics container ready"
exec "$@"
Build the container remotely:
apptainer build biotools.sif biotools.def
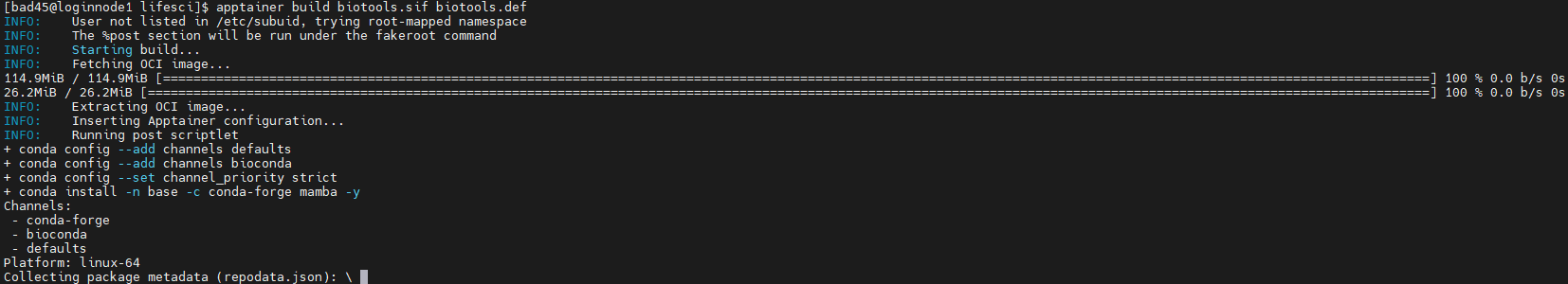
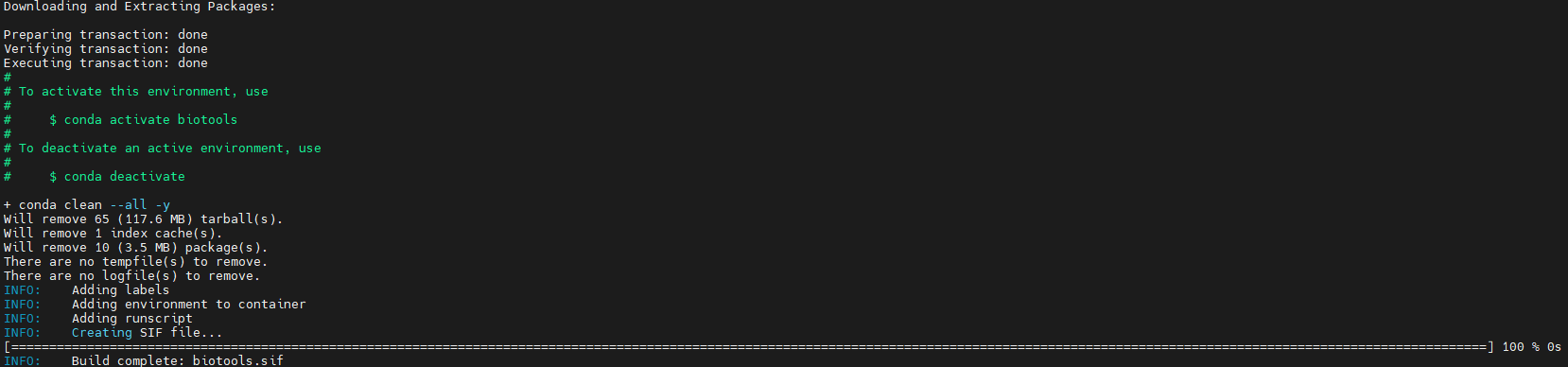
Option 2: Build Locally and Transfer
If you have a local machine with sudo access:
# On your local machine
sudo apptainer build biotools.sif biotools.def
# Transfer to Anatra
scp biotools.sif username@anatra.bath.ac.uk:/lifesciences/your-project/containers/
Example Container Job Script¶
Below is an example batch script for running a containerized bioinformatics workflow on Anatra:
#!/bin/bash
#SBATCH --job-name=apptainer-bio
#SBATCH --partition=lifesci # Use appropriate partition
#SBATCH --ntasks=1
#SBATCH --cpus-per-task=8
#SBATCH --mem=32G
#SBATCH --time=04:00:00
#SBATCH --output=apptainer_%j.out
# Enable detailed error reporting
set -eu
# Load Apptainer module
module purge
module load apptainer/1.4.1
# Set paths
CONTAINER="/lifesciences/containers/<sif file>"
INPUT_DIR="/lifesciences/your-project/data"
OUTPUT_DIR="/lifesciences/your-project/results"
INPUT_FILE=$1
# Create output directory if it doesn't exist
mkdir -p "$OUTPUT_DIR"
# Run Application
echo "Running on $INPUT_FILE..."
apptainer exec "$CONTAINER" "$INPUT_DIR/$INPUT_FILE"
echo "Analysis complete. Results in $OUTPUT_DIR"
Save this as run_script.slm and submit with:
sbatch run_script.slm your_file
Checking Container Information¶
# View container metadata
apptainer inspect biotools.sif
# See the definition file used to build it
apptainer inspect --deffile biotools.sif
# List available apps (if any)
apptainer inspect --list-apps biotools.sif
# Check what's in the environment
apptainer exec biotools.sif conda list